Pistoia Alliance US Conference 2015 - 1.5.4 New data - Nikolaus Schultz
- 1. Nikolaus Schultz Marie Josée and Henry R. Kravis Center for Molecular Oncology Memorial Sloan Kettering Cancer Center October 13, 2015 Visualization and Analysis of Cancer Genomics Data
- 2. Cost of DNA Sequencing is dropping rapidly
- 3. The Hallmarks of Cancer Hanahan and Weinberg. Cell. March 4 2011. Cancer is a class of diseases in which a group of cells display: uncontrolled growth invasion that intrudes upon and destroys adjacent tissues, and sometimes metastasis (spreading to other locations in the body via lymph or blood)
- 4. Many of these mechanisms are known, many not. Only some are treatable. All these properties are caused by genetic or epigenetic alterations. Can we identify the responsible alterations in the genomes of cancer patients? The Hallmarks of Cancer Cancer is a class of diseases in which a group of cells display: uncontrolled growth invasion that intrudes upon and destroys adjacent tissues, and sometimes metastasis (spreading to other locations in the body via lymph or blood)
- 5. Tumor development / Drivers versus passengers How does a cancer cell acquire all these different alterations? Sequential accumulation of genomic alterations that confer a growth advantage (like in evolution, but faster). Certain early events can increase the rate of accumulation, like mutations in DNA damage repair genes or cell-cycle checkpoint genes (or mutagens). Over time, many alterations develop. The ones that confer a growth advantage are called “drivers”, all others are “passengers”. Can we distinguish between them?
- 6. Identification of functional alterations in genomic data - per disease - per gene - per patient - per pathway Different, recurrent ways to alter the same pathway / process? Many events are rare, so we need hundreds of samples of the same disease (sub-)type to find them based on recurrence! Clinical applications: Development of new prognostic tools Identification of new treatment options Patient-specific treatment Utility of cancer genomics data bioinformaticians biologists clinicians
- 7. 2010 2011 2012 2013 Kidney clear cell Endometrial cancer Thyroid cancer Head & neck squamous Lung squamous cell carcinoma Colorectal cancer Breast cancer Low grade glioma 2014 GBM Phase II Bladder cancer Lung adenocarcinoma Melanoma Prostate cancer Stomach adenocarcinoma + lobular breast cancer, chromophobe kidney, papillary kidney,, pancreatic, rare tumors … 500 samples per tumor type 10,000 tumor / normal pairs total The Cancer Genome Atlas Project History 20092008 Ovarian cancer GBM AML Cervical Liver Sarcoma 2015
- 8. Cancer Cell Line Encyclopedia (CCLE) Broad Institute, Sanger, Washington University, etc. Tumor sequencing in hospitals (MSKCC 500 per month) Sources of tumor sequencing data 10,000 tumors 6,000 tumors 1,000 cell lines 5,000 tumors >15,000 tumors
- 9. Raw data (FASTQ / BAM files) dbGaP, CGHub, ICGC Data Portal Processed data (gene level data, mutation calls) TCGA Data Portal, ICGC Data Portal, SupplementaryTables Data slices (subsets of processed data) Data visualization and analysis tools Data availability bioinformaticians biologists, clinicians
- 10. Raw data (FASTQ / BAM files) dbGaP, CGHub, ICGC Data Portal Processed data (gene level data, mutation calls) TCGA Data Portal, ICGC Data Portal, SupplementaryTables Data slices (subsets of processed data) Data visualization and analysis tools Data availability bioinformaticians biologists, clinicians Reduction of complexity!
- 11. Most mutations found in cancer are “passengers”
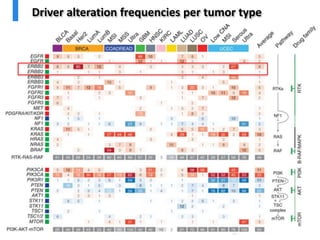
- 13. Driver alteration frequencies per tumor type
- 14. Driver alteration frequencies per tumor type
- 15. Rec L domain Furin-like Rec L domain Kinase domain ERBB2 mutation hotspots across cancer types
- 16. Rec L domain Furin-like Rec L domain Kinase domain ERBB2 mutation hotspots across cancer types signal noise
- 17. ERBB2 mutation hotspots across cancer types S310F Bladder: 1 Breast: 3 Cervical: 1 Colorectal: 2 Lung adeno: 2 Ovarian: 2 Stomach: 1 CCLE: 1 (bladder) L755S/M/P/W Breast: 4 Colorectal: 2 Endometrial: 1 Kidney (pap): 1 Melanoma: 1 Stomach: 1 CCLE: 3 (colorectal, stomach, brain) V777L/A Breast: 1 Colorectal: 2 GBM: 2 V842I Breast: 1 Colorectal: 4 Endometrial: 2 CCLE: 4 (Lung, ovarian, endometrial) R678Q Breast: 1 Colorectal: 1 Endometrial: 1 Stomach: 2 CCLE: 1 (colorectal) 774-776ins Lung adeno: 6 CCLE: 1 (lung) Rec L domain Furin-like Rec L domain Kinase domain
- 18. ERBB2 mutation hotspots across cancer types S310F Bladder: 1 Breast: 3 Cervical: 1 Colorectal: 2 Lung adeno: 2 Ovarian: 2 Stomach: 1 CCLE: 1 (bladder) L755S/M/P/W Breast: 4 Colorectal: 2 Endometrial: 1 Kidney (pap): 1 Melanoma: 1 Stomach: 1 CCLE: 3 (colorectal, stomach, brain) V777L/A Breast: 1 Colorectal: 2 GBM: 2 V842I Breast: 1 Colorectal: 4 Endometrial: 2 CCLE: 4 (Lung, ovarian, endometrial) R678Q Breast: 1 Colorectal: 1 Endometrial: 1 Stomach: 2 CCLE: 1 (colorectal) 774-776ins Lung adeno: 6 CCLE: 1 (lung) Rec L domain Furin-like Rec L domain Kinase domain Greulich et al. PNAS 2012. Kancha et al. PLoS ONE 2011. Bose et al. Cancer Discovery 2012. Bose et al. Cancer Discovery 2012. Bose et al. Cancer Discovery 2012.
- 19. cBioPortal for Cancer Genomics: Data to knowledge Tumor DNA DNA sequencer, microarrays … Tumor and normal sequences Data Intuitive interface, quick response time, reduction of complexity
- 20. Alteration types and thresholds can be customized for each gene. Reduction of complexity: Event calls Which genes are altered in which samples?
- 21. cBioPortal Data visualization and exploration in cBioPortal Clinical MSK-IMPACT Genomicdata CMO Research Foundation Medicine
- 22. Clinical MSK-IMPACT Genomicdata CMO Research cBioPortal Data visualization and exploration in cBioPortal TCGA, ICGC Other public data Foundation Medicine
- 23. Clinical MSK-IMPACT Genomicdata CMO Research cBioPortal Foundation Medicine TCGA, ICGC Other public data MSKCC clinical data Data visualization and exploration in cBioPortal
- 24. Clinical MSK-IMPACT Genomicdata CMO Research cBioPortal OncoKB: Annotation of variant effects, treatment Foundation Medicine TCGA, ICGC Other public data Clinical annotation Step 1: Manual Step 2: Automated via institutional databases MSKCC clinical data OncoKB Knowledgebase of oncogenic mutations Variant effect NCCN guidelines Standard therapy Investigational therapy Clinical trials
- 25. Live demonstration of cBioPortal http://cbioportal.org/
- 26. cBioPortal usage and interest cbioportal.org >5,000 unique users per week, doubling every year
- 27. cBioPortal usage and interest cbioportal.org >5,000 unique users per week, doubling every year Numerous academic installations of cBioPortal: Dana-Farber, Princess Margaret, CHOP, Weill Cornell, Fred Hutchinson, UCSC, Columbia, NYU, NY Genome Center, British Columbia, University of Michigan, SickKids, Vanderbilt, Emory, UNC, University of Pittsburgh, CRUK, EMBL, Charite Berlin, institutions in Japan, China, … Interest by several people to modify or customize the code, and to contribute new features Interest by pharmaceutical companies and others to use cBioPortal ● For internal data analysis (large pharma) ● In customer-facing applications (smaller service companies)
- 28. Switch to open source cBioPortal source code is available via GitHub: https://github.com/cBioPortal/cbioportal AGPL license v3 (Affero GPL): A GPL variant, main difference is that redistribution over a network triggers the copyleft requirements Impact on cancer research, patient treatment, drug development through: • More robust and flexible software • Accelerated development of new features • Wider user base, collaborative culture
- 29. Core cBioPortal Development group Memorial Sloan Kettering Cancer Center Nikolaus Schultz, Chris Sander, Benjamin Gross, JJ Gao Dana Farber Cancer Institute Ethan Cerami Princess Margaret Cancer Centre Trevor Pugh, Stuart Watt Re-uniting two cBioPortal founders Coordination of architectural decisions, feature development, merges, etc. TheHyve now offering commercial services around cBioPortal
- 30. Summary Rapidly growing body of cancer genomics data (public and private) Reduction of complexity can make these data accessible and interpretable cBioPortal allows access to cancer genomics data sets: cbioportal.org: public site via GitHub: install local versions cBioPortal is now fully open source software data pipelines and data sets coming soon commercial support available Still exploring pre-competitive funding options
- 31. Acknowledgements CMO Cyriac Kandoth William Lee Rajmohan Murali Nicholas D. Socci BarryTaylor Michael Berger Agnes Viale David B. Solit MichaelTrapani Ederlinda Paraiso Molecular Diagnostics Ahmet Zehir Aijaz Syed Donavan Cheng Michael Berger Maria Arcila Marc Ladanyi Information Systems Mike Eubanks Stu Gardos cBioPortal JianJiong Gao Benjamin Gross Yichao Sun Hongxin Zhang Fred Criscuolo Dong Li Adam Abeshouse Ritika Kundra Annice Chen Chris Sander Onur Sumer Arman Aksoy Ethan Cerami Knowledgebase Debyani Chakravarty Sarah Phillips Julia Rudolph Bioinformatics Core Joanne Edington
- 32. Demo slides
- 53. End of Live Demo